
Concept explainers
Common baker’s yeast (
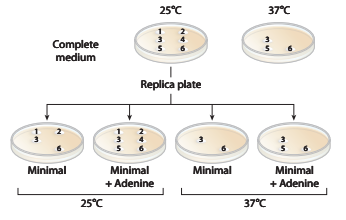
a. Which colonies are prototrophic and which are auxotrophic? What growth information is used to makethese determinations?
b. Classify the nature of the mutations in colonies
c. What can you say about colony
Want to see the full answer?
Check out a sample textbook solution
Chapter 11 Solutions
Pearson eText Genetic Analysis: An Integrated Approach -- Instant Access (Pearson+)
- Assume you have successfully cloned a small (200 bp) fragment of DNA into the polylinker region of a pUC18 cloning vector. Describe the appearance of transformed colonies you would expect to see on each of the following plates: plain media, media containing ampicillin, media containing tetracycline, media containing ampicillin and X-Gal.arrow_forwardpZERO®-1 is a 2808 bp cloning vector from Invitrogen. This vector allows effective selection of positive recombinants via disruption of the lethal gene, ccdB. Besides the lethal gene, it has an additional Zeocin resistance gene for tighter screening control. The Zeocin resistance gene in this vector is certainly more advantageous over ampicillin resistance gene. Give its THREE (3) advantages.arrow_forwardThe following two strains of E. coli are crossed with each other: Hfr pan* thi* ala* and F¯pan¯ thi¯ ala¯ It was shown that the pan marker entered last in interrupted conjugation experiments. By spreading the bacteria from these experiments on different selection media, the following results (in number of colonies) were obtained: MM + Gluc IM + Gluc + ala MM + Gluc + thi MM + Gluc + thi + ala 280 281 286 339 Give the number of each phenotypic class and justify your answer. Determine the order of the genes. What are the genetic distances between the different loci? Abbreviations: Gluc: glucose; Ala: alanine; Thi: thiamine; Pan: pantothenic acid.arrow_forward
- A shuttle vector is a vector (usually a plasmid) constructed so that it can propagate in two different host species. One of the most common types of shuttle vectors is the yeast shuttle vector. Examples of such vectors derived from yeast are Yeast Episomal Plasmid (YEp), Yeast Integrating Plasmid (YIp) and Yeast Replicating Plasmid (YRp). Among these three vectors, YIp has the lowest transformation frequency and copy number per cell. Explain why Ylp is still popularly used despite its limitations.arrow_forwardDetermine the map distances between the genes.arrow_forwardMatch the following Conjugation terms with the most appropriate description in the image: F- recombinant F- strain F+ strain HFR strainarrow_forward
- Order the following experimental steps to identify auxotrophic mutants in Saccharomyces Cerevisiae that cannot sythesize their own Leucine. an option can be selected more than once. step 1: Step 2: Step 3: Step 4: Step 5: Step 6:arrow_forwardGiven what we've discussed in class, what will be most likely outcome if you conjugate an streptomycin resistant ampicillin sensitive methionine auxotroph E. coli strain (engineered to be pir+) that is F- with a streptomycin sensitive non-HFR methionine prototroph strain that is F- and RP4+ but contains pUC18? Colonies on minimal media + ampicillin +streptomycin plates No colonies on minimal media +ampicillin +streptomycin platesarrow_forwardIn E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What information do you know based onthe question and your understanding of the topic?arrow_forward
- In E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What topic in genetics does this question address?arrow_forwardA fluctuation test was carried out to determine the rate of mutation to Azetidine resistance (a toxic proline analog) in S. typhimurium. Twenty tubes of rich medium were each inoculated with a few wild-type cells and the cultures were grown to 10° cells / ml. A 0.1 ml sample of each culture was then plated on each plate (a total of 20 plates) to detect AztR mutants. The results are shown in the following table. Calculate the mutation rate of S. typhimurium to AztR. Culture # # AztR mutants Culture # # AztR mutants 11 12 3 4. 13 14 15 4. 30 303 97 69 14 16 17 18 19 20 10 19arrow_forwardFrom an Escherichia coli strain, five Hfr strains were isolated. The location and orientation of the transfer origin of each Hfr strain is shown in Figure 1. You want to use these five strains to map the locus responsible for thiamine synthesis, called thi. Each Hfr strain is sensitive to rifampicin (Rifs) and thi*. Conjugation experiments are performed between each of the Hfr strains and an F strain Rif Thi™. 0 T leu 10 20 T nadD pyrC trp 40 T his 60 70 2) The results are shown in the following table: Donor strain Hfr1 Hfr2 Hfr3 Hfr4 Hfr5 cysG 80 90 1) What is the selection medium used in these conjugation experiments metA Colonies Thi+ 1000 0 400 0 25 100 Hfr1 Hfr2 Figure 1: Chromosome map of Escherichia coli. Five Hfr strains (Hfr1 to Hfr5) were isolated and the location and orientation of the origin of transfer is shown by the arrows in each Hfr strain. Distances in minutes are shown. Leu: leucine biosynthesis; nadD: NAD biosynthesis; pyrC: pyrimidine biosynthesis; trp: tryptophan…arrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
