
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 14, Problem 29P
Streptococcus parasanguis is a bacterial species that initiates dental plaque formation by adhering to teeth. To investigate ways to eliminate plaque, researchers constructed a plasmid, depicted in the figure shown, to mutagenize S. parasanguis. The key features of this plasmid include repts A (a temperature-sensitive origin of replication), kanr (a gene for resistance to the antibiotic kanamycin), and the transposon IS256. This transposon contains the erm gene for resistance to the antibiotic erythromycin. IS256 transposes in S. parasanguis thanks to a gene encoding a transposase enzyme that moves all DNA sequences located between the transposon’s inverted repeats (IRs).
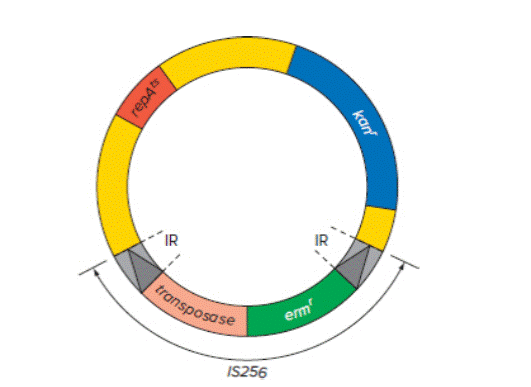
| a. | How could the researchers use this plasmid as a mutagen? Consider how they could get the transposon into the bacteria, and how they could identify strains that had new insertions of IS256 into S. parasanguis genes. Your answer should explain why the plasmid has two different antibiotic resistance genes as well as a temperature-sensitive origin of replication. |
| b. | Why would the researchers use this plasmid as a mutagen? |
| c. | If the investigators found a mutant strain of S. parasanguis that was defective in plaque formation, how could they identify the affected gene? |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
If I clone a complete eukaryotic gene, including the eukaryotic promoter region, ligate it into a plasmid, and transform it into E. coli, will I be able use the transformed E. coli to make the corresponding protein? Explain why, or why not? If you decide to do this, what would your cloning strategy be?
The gene atg-12 codes for a protein associated with abnormal rates of cell destruction and recycling of cell contents. Scientists studying bacterial plasmids devised an experiment using recombinant DNA techniques to remove a section of DNA (gene atg-12) of a bacterial plasmid (pOKE103) and create a new plasmid (pOKE104) that did not contain the gene atg-12. The new plasmid was then incorporated within the DNA of the fungus Neurospora crassa in cellular studies.
Which statement explains the expected heredity of fungi that incorporate pOKE104?
A - Fungi that incorporate pOKE104 will produce the protein from gene atg-12 and have increased rates of malignant tumors.
B - Fungi that incorporate pOKE104 will not produce the protein from gene atg-12 but will have a normal appearance.
C - The prokaryotic DNA will remain separate within the fungal cells, produce the protein from gene atg-12, and have increased rates of malignant tumors.
D - The prokaryotic DNA will remain separate within the…
A 2.0kb bacterial plasmid ‘BS1030’ is digested with the restriction endonuclease Sau3A; the plasmid map is depicted in the diagram below and the Sau3A (S) restriction sites are indicated.
Which of the following DNA fragments do you expect to see on an agarose gel when you run Sau3A-digested plasmid ‘BS1030’ DNA?
a.
250 bp, 450 bp, 550 bp, 1.1 kb, 1.5 kb and 2.0 kb
b.
2.0kb
c.
250 bp, 400 bp, 450 bp, 500 bp and 550 bp
d.
100 bp, 200 bp, 250 bp, 400 bp, 500 bp and 550 bp
Chapter 14 Solutions
Genetics: From Genes to Genomes
Ch. 14 - Choose the phrase from the right column that best...Ch. 14 - The unicellular, rod-shaped bacterium E. coli is 2...Ch. 14 - Now that the sequence of the entire E. coli K12...Ch. 14 - Bacterial genomes such as that of E. coli...Ch. 14 - List at least three features of eukaryotic genomes...Ch. 14 - Describe a mechanism by which a gene could move...Ch. 14 - High salt concentrations tend to cause protein...Ch. 14 - Recently, scientists tested the possibility that...Ch. 14 - A recent metagenomic study analyzed the...Ch. 14 - Linezolid is a new type of antibiotic that...
Ch. 14 - A liquid culture of E. coli at a concentration of...Ch. 14 - Pick out the medium i, ii, iii, or iv onto which...Ch. 14 - This problem concerns Fig. 14.14, which...Ch. 14 - In two isolates one is resistant to ampicillin,...Ch. 14 - E. coli cells usually have only one copy of the F...Ch. 14 - In E. coli, the genes purC and pyrB are located...Ch. 14 - DNA sequencing of the entire H. influenzae genome...Ch. 14 - Genes encoding toxins are often located on...Ch. 14 - a. You want to perform an interrupted-mating...Ch. 14 - In Problem 19, do you think that most of the...Ch. 14 - One issue with interrupted-mating experiments such...Ch. 14 - Prob. 22PCh. 14 - Starting with an F- strain that was prototrophic...Ch. 14 - You can carry out matings between an Hfr and F...Ch. 14 - Genome sequences show that some pathogenic...Ch. 14 - Generalized and specialized transduction both...Ch. 14 - This problem highlights some useful variations of...Ch. 14 - A researcher has a Trp auxotrophic strain of E....Ch. 14 - Streptococcus parasanguis is a bacterial species...Ch. 14 - The sequence at one end of one strand of the...Ch. 14 - Scientists who study amino acid biosynthesis...Ch. 14 - Suppose that you could obtain radioactively...Ch. 14 - Prob. 34PCh. 14 - Some scientists are trying to engineer...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- In producing genetically engineered human insulin in bacteria, why is it important to use the samerestriction enzyme to cut both the human DNA and the bacterial plasmid?arrow_forwardA PCR reaction was performed to amplify the XULA3 gene, which is bp 882-5,364 on a plasmid that is 11,719 bp. After the PCR, the product was digested with XhoI. There are XhoI sites on the plasmid at bp 1,434, 4,655, and 7,368. Calculate the size(s) that would result when the product is digested with XhoI. Then enter the size of the largest fragment (in bp).arrow_forwardBelow is a map of the E. coli cloning vector after the insertion of pka-1. Ori denotes the origin of replication; amp denotes the ampicillin resistance gene. HindIII, BamHI, SalI, and EcoRI designate restriction enzyme sites. There are no other restriction enzyme sites found on this vector. The numerals denote the number of base pairs between different locations on the plasmid. For instance, there are 400 base pairs between the HindIII and BamHI site, and there are 3000 base pairs in the entire cloning vector (following the integration of pka-1). With the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following: Isolation of target DNA fragments (often referred to as inserts) Ligation of inserts into the plasmid, creating recombinant molecules Transformation of recombinant plasmids into bacteria or other suitable host for propagation Screening/selection of hosts containing the intended…arrow_forward
- In cloning human DNA, why is it necessary to insert the DNA into a vector such as a bacterial plasmid?arrow_forwardConstruct a restriction map for Plasmid X to show the recognition sites for two commonly used restriction enzymes: EcoRI and BamHI. Plasmid X undigested Plasmid X digested with EcoRI Plasmid X digested with BamHI Plasmid X digested with EcoRI and BamHI 2200 bp 1000 bp 1200 bp 400 bp 1800 bp 200 bp 400 bp 600 bp 1000 bparrow_forward. For Figure 15-7, draw out a series of steps that could explain the origin of this large plasmid containing manytransposable elements.arrow_forward
- This plasmid was digested using different restriction enzymes whose sites have been mapped. The plasmid is 7896 base pairs long. This is a long question so u can count this as two or even three but please answer the question? Determine the size (base pairs) and number of fragments that would be produced if the plasmid was digested with the following enzymes: a) EcoRI b) BamHI c) HindIII d) EcoRI and HindIII e)EcoRI, HindIII, and BamHI *Hint- this is actually an EASY question, since the restriction map is already drawn for you!arrow_forwardKnowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardIn the formation of recombinant DNA, a restriction endonuclease cuts a bacterial plasmid to give sticky ends. The DNA segments that are to be added to the plasmid are cleaved with the same restriction endonuclease. What aresticky ends and why is it important that the target DNA and the plasmid it will be incorporated into have complementary sticky ends?arrow_forward
- Consider the ends of the DNA fragments shown below. They have been produced by digestion of a single sequence of DNA using a number of restriction endonucleases. 1. 5'A 3' 3'TTCGA5' 2. 5'G 3' 3'CAGCT5' 3. 5'AATTC3' 3' G5 4. 5'TCGAC3' 3' G5' 5. 5'GGG 3' 3'CCC 5' Which of these ends are capable of annealing and being joined by DNA ligase?arrow_forwardOne would be correct in assuming that the restriction sites present in the multiplecloning site of the pUC18 plasmid willarrow_forwardIn relation to the use of restriction enzymes in recombinant DNA technology, answer the following: You have accidentally torn the labels off two tubes (tube A and tube B), each containing a different plasmid, now you do not know which plasmid is in which tube. Fortunately, you have restriction maps for both plasmids, shown in Figure below. You have the opportunity to test just one sample from one of your tubes. By utilizing agarose gel electrophoresis technique, which restriction enzyme OR combination of restriction enzymes would you use in this experiment to determine which plasmid is found in which tube?. (Hint: if you use Hind III restriction enzyme you are going to get ONE single fragment with a molecular size of → 0.5+0.3+0.2+0.4+1+1 = 3.4 kb).arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license