
Concept explainers
An electrophoretic mobility shift assay can be used to study the binding of proteins to a segment of DNA. In the experiment shown here, an EMSA was used to examine the requirements for the binding of RNA polymerase II (from eukaryotic cells) to the promoter of a protein-encoding gene. The assembly of general transcription factors and RNA polymerase II at the core promoter is described in Chapter 12 (Figure 12.14). In this experiment, the segment of DNA containing a promoter sequence was 1100 bp in length. The fragment was mixed with various combinations of proteins and then subjected to an EMSA.
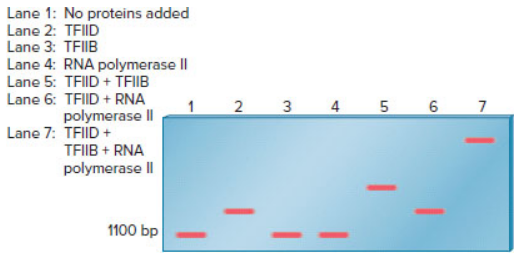
Explain which proteins (TFIID, TFIIB, or RNA polymerase II) are able to bind to this DNA fragment by themselves. Which transcription factors (i.e., TFIID or TFIIB) are needed for the binding of RNA polymerase II?
Want to see the full answer?
Check out a sample textbook solution
Chapter 21 Solutions
Genetics: Analysis and Principles
- When a region of DNA that contains the genetic information for a protein is isolated from a bacterial cell and inserted into a eukaryotic cell in a proper position between a promoter and a terminator, the resulting cell usually produces the correct protein. But when the experiment is done in the reverse direction (eukaryotic DNA into a bacterial cell), the correct protein is often not produced. Can you suggest an explanation?arrow_forwardA mutant strain of Salmonella bacteria carries a mutation of the rho protein that has fully activity at 37°C but is completely inactivated when the mutant strain is grown at 40°C. a)Speculate about the kind of differences you would expect to see if you compared a broad spectrum of mRNAs from the mutant strain grown at 37°C and the same spectrum of mRNAs from the strain when grown at 40°C. b)Are all the mRNAs affected by the rho protein mutation in the same way? Why or why not?arrow_forwardHow do you think that transcription randomizes positions of nucleosomes and repression restores the ordering after transcription? How might you test to see if there was an exchange of histone subunits during transcription or if the nucleosome is truly transferred as a single unit? Would you expect the DNA band representing the distance from the restriction enzyme site to the hypersensitive site to be a single band or a smear? Defend your answer.arrow_forward
- Upon identification of the DNA regulatory sequence responsible for translating a given gene, you note that it is enriched with CG sequences. Is the corresponding gene likely to be a highly expressed transcript?arrow_forwardMutations in bacterial promoters may increase or decrease the rate of gene transcription. Promoter mutations that increase the transcription rate are termed up-promoter mutations, and those that decrease the transcription rate are termed down-promoter mutations. As shown , the sequence of the −10 site of the promoter for the lac operon is TATGTT. Would you expect the following mutations to be up-promoter or down-promoter mutations? A. TATGTT to TATATT B. TATGTT to TTTGTT C. TATGTT to TATGATarrow_forwardSuppose you want to study the transcription in vitro of one particular gene in a DNA molecule that contains several genes and promoters. Without adding specific regulatory proteins, how might you stimulate transcription from the gene of interest relative to the transcription of the other genes on your DNA template? To make all of the complexes identical, you would like to arrest all transcriptional events at the same position on the DNA template before isolating the complex. How might you do this?arrow_forward
- Consider the Rho-dependent terminator sequence 5’CCCAGCCCGCCUAAUGAGCGGCCUUUUUUUU-3’. What affect would a point mutation at any one of the bolded and underlined nucleotides disrupt termination of transcription? Group of answer choices Mutation in one of these nucleotides would disrupt base pairing, preventing the formation of the hairpin and disrupting termination. Mutation in one of these nucleotides would have no affect on base pairing, so the termination hairpin is formed and termination proceeds. Mutation in one of these nucleotides would not disrupt base pairing, but would prevent the formation of the hairpin and disrupt termination. Mutation in one of these nucleotides would disrupt base pairing, but not affect the formation of the hairpin and termination proceeds.arrow_forwardIf the lacZ protein breaks down lactose, is it worthwhile to make it when there is no lactose around? How does the bacteria use this system to efficiently control the production of the lacZ protein? Does the presence of lactose in the cell alter its ability to repress translation? To what does the lacI protein bind to? What effect does the lacI gene have on transcription of the lacZ and lacY genes?arrow_forward. One mechanism by which antisense RNAs act as negative regulators of gene expression is by base pairingwith the ribosome binding site on the sense mRNA toblock translation. In a second, alternative mechanism,the act of transcribing an antisense RNA can somehow prevent RNA polymerase from recognizing thesense promoter for the same gene. Design an experimental approach that would enable you to distinguishbetween these two modes of action at a specific gene.(Hint: What would be the outcome in each case ifhigh levels of the antisense RNA were transcribedfrom a gene on a plasmid?)arrow_forward
- Gal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forwardMany eukaryotic promoter regions contain CAAT boxes with consensus sequences CAAT or CCAAT approximately 70 to 80 bases upstream from the transcription start site. How might one determine the influence of CAAT boxes on the transcription rate of a given gene?arrow_forwardMutations in the promoter region of the B-globin gene indicate that some areas are more sensitive than others. When mutations occur in consensus sequences(modular elements such as GC box, CAAT box, and TATA box), does transcription usually increase or decrease? Explain.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





