
wo techniques commonly used to study the expression patterns of genes that play a role in development are Northern blotting and in situ hybridization. As described in Chapter 21, Northern blotting is used to detect RNA that is transcribed from a particular gene. In this method, a specific RNA is detected by using a short segment of cloned DNA as a probe. The DNA probe, which is labeled, is complementary to the RNA that the researcher wishes to detect. After the DNA probe binds to the RNA within a blot of a gel, the RNA is visualized as a labeled band on a nylon membrane. For example, a DNA probe that is complementary to the bicoid mRNA could be used to specifically detect the amount of that mRNA in a blot.
A second technique, termed fluorescence in situ hybridization (FISH), is used to identify the locations of genes on chromosomes. This technique is also used to locate gene products within oocytes, embryos, and larvae. Thus, it has been commonly used by developmental geneticists to understand the expression patterns of genes during development. The micrograph in Figure 26.8b is derived from the application of the FISH technique. In this case, the probe was complementary to bicoid mRNA.
Now here is the question. Suppose a researcher has three different Drosophila strains that have loss-of-function mutations in the bicoid gene. We will call them bicoid-A, bicoid-B, and bicoid-C; the wild type is designated bicoid+. To study these mutations,
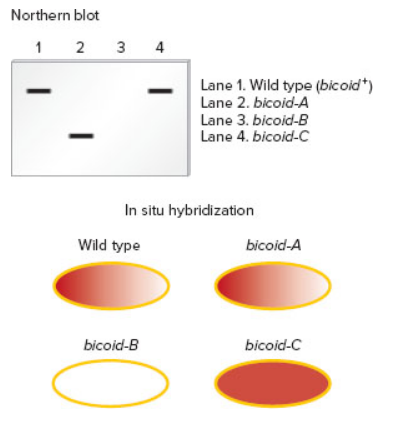
A. How can phenotypically normal female flies be homozygous for a loss-of-function allele in the bicoid gene?
B. Explain the type of mutation (e.g., deletion, point mutation, etc.) in each of the three strains. Explain how the mutation may cause a loss of normal function for the bicoid gene product.
C. Discuss how the use of both techniques provides more definitive information than the application of just one of the techniques.
Want to see the full answer?
Check out a sample textbook solution
Chapter 26 Solutions
Genetics: Analysis and Principles
- Scientists carried out a microarray analysis to compare the gene expression of normal pancreatic cells to that of cancer cells from a person with pancreatic cancer. The scientists labeled the cDNA from the normal pancreatic cells with green fluorescent nucleotides. They labeled the cDNA from the cancer cells with red fluorescent nucleotides. The two cDNAs were mixed and allowed to hybridize to a microarray. Less p53 activity is found in cancer pancreatic cells than normal cells. What color would the spot for the p53 gene be on the microarray? Red Green Yellow Blackarrow_forwardA GWAS study using genomic resequencing may find a statistically significant SNP that is: (choose all that apply) Group of answer choices near a regulatory sequence that causes a change in a gene's expression resulting in the phenotype studied. in a gene's exon that changes that gene's product and produces the phenotype studied. near a gene that causes the phenotype studied. in a regulatory sequence that changes the expression of a gene resulting in the phenotype studied.arrow_forwardProtein levels and mRNA levels for a particualr gene don’t always match. For example, the GCN4 gene in yeast is always producing mRNA, but the Gcn4 protein is only made when the cells are starved. B. What does this mean for diagnostic techniques that try to look at gene expression?arrow_forward
- In a particular organism, there are two similar genes called YFG1 and YFG2. YFG1 is expressed in the liver and not in the pancreas, and YFG2 is expressed in the pancreas but not the liver. Neither YFG1 nor YFG2 is expressed in the heart. If you extract DNA from heart cells, do you expect to see the YFG2 gene? Explain why. Do you expect to see the YFG1 protein when you analyze protein extract from liver cells? And from pancreas cells? And from heart cells? Explain why. Is it possible to produce YFG1 and YFG2 proteins via alternative splicing? Explain one possible way (mechanism) to regulate the expression of YFG1 gene.arrow_forwardWhat does it mean for a transposable element to be effectively “dead”? A. The transposable elements are “dead” because they are only found in non-coding regions and therefore do not interfere with phenotypic expression. B. The transposable elements are “dead” because they are no longer able to undergo transposition and move to another region of the genome. C. The transposable elements are “dead” because they do cause disease despite their presence. D. The transposable elements are “dead” because they occur only in somatic cells and therefore are not heritable.arrow_forwardwhich is the following is not a step in the process of de novo gene birth? o translated peptides provide some benefit to the orhganism, leading to selection for increased expression. o the open reading frame picks up sequences that enable recognition of its transcribed mRNA by the ribosome. o an open reading frame is created through mutations. o the gene copy number changes as a result of gene duplication to make paralogs.arrow_forward
- Protein levels and mRNA levels for a particualr gene don’t always match. For example, the GCN4 gene in yeast is always producing mRNA, but the Gcn4 protein is only made when the cells are starved. What does this mean for diagnostic techniques that try to look at gene expression?arrow_forwardDraw a basket mutant embryo. What does basket encode? Why do the mutant embryos have this phenotype?arrow_forwardAn enhancer, located upstream from a gene, has the following sequence: 5′–GTAG–3′ 3′–CATC–5′ This enhancer is orientation-independent. Which of the following sequences also works as an enhancer? A. 5′–CTAC–3′ 3′–GATG–5′ B. 5′–GATG–3′ 3′–CTAC–5′ C. 5′–CATC–3′ 3′–GTAG–5′ C15.arrow_forward
- TOH1 protein is typically expressed at the same level in kidney and stomach cells and is essential to appropriate function of both organs. TOH disorder is a disorder where no detectable TOH1 protein is present in kidney cells due to a mutation in the promoter leading to no transcription. No other organs are affected in TOH disorder. You used microarray analysis to examine cells from kidney and stomach cells from the same individual with TOH disorder. You labeled the cDNA from the affected kidney cells with red fluorescent nucleotides, and you labeled the cDNA from the unaffected stomach cells with green fluorescent nucleotides. After you mixed the cDNAs and allowed for hybridization, what color would you expect to see for the spot for the TOH1 gene when you analyze the microarray data? a) Red b) Green c) Black d) Yellowarrow_forwardThe insertion of transposable elements into genes can alter the normal pattern of expression. In the following situations, describe the possible consequences on gene expression.a. A LINE inserts into an enhancer of a human gene. b. A transposable element contains a binding site for a transcriptional repressor and inserts adjacent to a promoter. c. An Alu element inserts into the 3′ splice (AG) site of an intron in a human gene. d. A Ds element that was inserted into the exon of a maize gene excises imperfectly and leaves three base pairs behind in the exon. e. Another excision by that same Ds element leaves two base pairs behind in the exon. f. A Ds element that was inserted into the middle of an intron excises imperfectly and leaves five base pairs behind in the intron.arrow_forwardWhy are some genes expressed and some not? Please be as detailed as possible.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





