
BIOCHEMISTRY W/1 TERM ACHEIVE ACCESS
9th Edition
ISBN: 9781319425746
Author: BERG
Publisher: MAC HIGHER
expand_more
expand_more
format_list_bulleted
Concept explainers
Question
Chapter 4, Problem 49P
Interpretation Introduction
Interpretation:
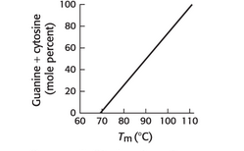
A reasonable explanation for the relation between % of GC base pairs in DNA and the melting temperature needs to be given.
Concept introduction:
The GC base pair i.e. Guanine - Cytosine pair help the DNA to maintain its helical structure which is dependent on the
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
What is the melting temp. of the following double-stranded DNA fragment
TCAAAAATCGAATATTTGCTTATCTA
AGTTTTTAGCTTATAAACGAATAGAT
Given this sequence ATGCACCG
About how often (every _____ bases) would this occur in double strand DNA that is known to contain 20%T. Show work.
Compare the melting temperature of a 1-kb segment of DNA containing20% A residues to that of a 1-kb segment containing 30% A residues under the same conditions.
Chapter 4 Solutions
BIOCHEMISTRY W/1 TERM ACHEIVE ACCESS
Ch. 4 - Prob. 1PCh. 4 - Prob. 2PCh. 4 - Prob. 3PCh. 4 - Prob. 4PCh. 4 - Prob. 5PCh. 4 - Prob. 6PCh. 4 - Prob. 7PCh. 4 - Prob. 8PCh. 4 - Prob. 9PCh. 4 - Prob. 10P
Ch. 4 - Prob. 11PCh. 4 - Prob. 12PCh. 4 - Prob. 13PCh. 4 - Prob. 14PCh. 4 - Prob. 15PCh. 4 - Prob. 16PCh. 4 - Prob. 17PCh. 4 - Prob. 18PCh. 4 - Prob. 19PCh. 4 - Prob. 20PCh. 4 - Prob. 21PCh. 4 - Prob. 22PCh. 4 - Prob. 23PCh. 4 - Prob. 24PCh. 4 - Prob. 25PCh. 4 - Prob. 26PCh. 4 - Prob. 27PCh. 4 - Prob. 28PCh. 4 - Prob. 29PCh. 4 - Prob. 30PCh. 4 - Prob. 31PCh. 4 - Prob. 32PCh. 4 - Prob. 33PCh. 4 - Prob. 34PCh. 4 - Prob. 35PCh. 4 - Prob. 36PCh. 4 - Prob. 37PCh. 4 - Prob. 38PCh. 4 - Prob. 39PCh. 4 - Prob. 40PCh. 4 - Prob. 41PCh. 4 - Prob. 42PCh. 4 - Prob. 43PCh. 4 - Prob. 44PCh. 4 - Prob. 45PCh. 4 - Prob. 46PCh. 4 - Prob. 47PCh. 4 - Prob. 48PCh. 4 - Prob. 49PCh. 4 - Prob. 50PCh. 4 - Prob. 51PCh. 4 - Prob. 52P
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- What is the melting temp. of the following double-stranded DNA fragment CATCGCGATCTGCAATTACGACGATAA GTAGCGCTAGACGTTAATGCTGCTATTarrow_forwardIt is known that double stranded DNA is denatured at low pH. pKa values should allow the determination of whether this is due to perturbation of the hydrogen bonding in A-T and/or G-C base pairs. The table gives values for the pKas of different protonated groups in the nucleobases.Nucleobase Position & pKa A N1, 3.5 G N7, 1.6; N1, 9.2 C N3, 4.2 T N3, 9.7a) Draw the A-T and G-C base pairs. - Label the bases with the one-letter code. – - Number the atoms in the rings and label the atom that attaches to the sugar. - Mark the groups that interact in normal…arrow_forwardHelicase Unwinding of the E. coli Chromosome Hexameric helicases, such as DnaB, the MCM proteins, and papilloma virus El helicase (illustrated in Figures 16.22 to 16.25), unwind DNA by passing one strand of the DNA duplex through the central pore, using a mechanism based on ATP-dependent binding interactions with the bases of that strand. The genome of E. coli K12 consists of 4,686,137 nucleotides. Assuming that DnaB functions like papilloma virus El helicase, from the information given in Chapter 16 on ATP-coupled DNA unwinding, calculate how many molecules of ATP would be needed to completely unwind the E. coli K 12 chromosome.arrow_forward
- Semiconservative or Conservative DNA Replication If 15N-Iabeled E. coli DNA has a density of 1.724 g/mL, 14N-labeled DNA has a density of 1.710 g/mL, and E. coli cells grown for many generations on 14NH4+as a nitrogen source are transferred to media containing 15NH4+as the sole N-source, (a) What will be the density of the DNA after one generation, assuming replication is semiconservative? (b) Suppose replication took place by a conservative mechanism in which the parental strands remained together and the two progeny strands were paired. Design an experiment that could distinguish between semiconservative and conservative modes of replication.arrow_forwardMay you please help me with this? A sample of purified DNA was incubated with deoxyribonuclease (DNAse) at 37oC. An aliquot was removed from the reaction mixture every minute for 5 minutes and the A260 recorded. The following data were obtained. Time (min) A260 0 0.60 1 0.64 2 0.67 3 0.70 4 0.72 5 0.73 Describe the action of deoxyribonuclease on DNA and explain the increase in A260.arrow_forwardRecall the DNA’s three-dimensional model. The DNA is a right-handed helix wherein onecomplete 360 0 turn covers a distance of 34 angstroms (Å) or 3.4 nm and 10 base pairs. As a result, thebase pairs are separated by a distance of approximately 3.4 Å. The diameter of the Watson and CrickDNA molecule is 20 Å.Calculate the average number of nucleotide pairs (or base pairs) per micrometer of DNA doublehelix according to the dimension mentioned above. Round off your answer to the nearest wholenumber. Note also that 1 micrometer = 10,000 angstroms.arrow_forward
- The Qiagen DNeasy DNA extraction kit is used to extract DNA from a liver cell line for subsequent molecular experiments. (i) (ii) for checking DNA purity using a spectrophotometer. Explain the principle behind this kit in DNA extraction. After the DNA is obtained, the purity of DNA should be determined. Discuss the stepsarrow_forwardThe complementary strands of DNA in the double helixare held together by hydrogen bonds: G ≡ C or A = T.These bonds can be broken (denatured) in aqueous solutions by heating to yield two single strands of DNA(see Figure 1-13a). How would you expect the relativeamounts of GC versus AT base pairs in a DNA doublehelix to affect the amount of heat required to denatureit? How would you expect the length of a DNA doublehelix in base pairs to affect the amount of heat requiredto denature it?arrow_forwardN. NH 2. One of the key pieces of information that Watson and Crick used in determining the secondary structure of DNA came from experiments done by E. Chargaff, in which he studied the nucleotide composition of DNA from many different species. O=P-OCH, N. `NH, HN он O= P- OCH, NH, Chargaff noted that the molar quantity of A_was always approximately equal to the molar quantity of T. and the molar quantity of C was always approximately equal to the molar quantity of G. How were Chargaff's results explained by the structural model of DNA proposed by Watson and Crick? N OH N. O= P-OCH, OH OHarrow_forward
- pppApCpCpUpApGpApU-OH(a) Using the straight-chain sugar convention, write the structure of the DNA strand that encoded this short stretch of RNA.(b) Using the simplest convention for representing the DNA base sequence, write the structure of the nontemplate DNA strand.arrow_forwardThe melting temperature Tm of DNA can be predicted by calculation without actually measuring it. Calculate the Tm of the DNA double strand shown in (1) to (3), and discuss the results. The numbers in parentheses indicate the degree of polymerization of nucleotides.(1) A(10) + T(10), (2) A(15) + T(15), (3) G(10) + C(10)arrow_forwardEstimate the viscosith of 1.0 vol % agarose gel solution if it took 58 minutes for 1500 base pairs long DNA molecule to migrate 11cm during gel electrophoresis under 110V. Assume appropriate numerical values for the missing parameters such as the charge and diameter of DNA molecules.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning

Biochemistry
Biochemistry
ISBN:9781305577206
Author:Reginald H. Garrett, Charles M. Grisham
Publisher:Cengage Learning
Macromolecules | Classes and Functions; Author: 2 Minute Classroom;https://www.youtube.com/watch?v=V5hhrDFo8Vk;License: Standard youtube license