
Concept explainers
DNA footprint protection (described in Research Technique
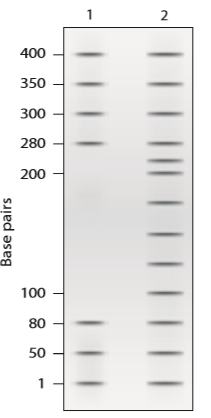
a. Explain why this gel provides evidence that the cloned DNA may act as a promoter sequence.
b. Approximately what length is the DNA region protected by RNA pol II and TFIIs?
c. What additional genetic experiments would you suggest to verify that this region of cloned DNA contains a functional promoter?
Want to see the full answer?
Check out a sample textbook solution
Chapter 8 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- How do you think that transcription randomizes positions of nucleosomes and repression restores the ordering after transcription? How might you test to see if there was an exchange of histone subunits during transcription or if the nucleosome is truly transferred as a single unit? Would you expect the DNA band representing the distance from the restriction enzyme site to the hypersensitive site to be a single band or a smear? Defend your answer.arrow_forwardThe interphase nucleus is a highly structured organelle with chromosome territories, interchromatin compartments, and transcription factories. In cultured human cells, researchers have identified approximately 8000 transcription factories per cell, each containing an average of eight tightly associated RNAP II molecules actively transcribing RNA. If each RNAP II molecule is transcribing a different gene, how might such a transcription factory appear? Provide a simple diagram that shows eight different genes being transcribed in a transcription factory and include the promoters, structural genes, and nascent transcripts in your presentation.arrow_forwardSuppose you want to study the transcription in vitro of one particular gene in a DNA molecule that contains several genes and promoters. Without adding specific regulatory proteins, how might you stimulate transcription from the gene of interest relative to the transcription of the other genes on your DNA template? To make all of the complexes identical, you would like to arrest all transcriptional events at the same position on the DNA template before isolating the complex. How might you do this?arrow_forward
- The following is a portion of an mRNA sequence: 3’ –AUCGUCAUGCAGA-5’ a)During transcription, was the adenine at the left-hand side of the sequence the first or the last nucleotide used to build the portion of the mRNA shown? Explain how you know. b)Write out the sequence and polarity of the DNA duplex that encodes this mRNA segment. Label the template and coding DNA strands. c)Identify the direction in which the promoter region for this gene will be located.arrow_forwardThe human hexokinase enzyme has the same function as the bacterial hexokinase enzyme but is somewhat different in its amino acid sequence. You have obtained a mutant bacterial strain in which the gene for hexokinase is missing. If you introduce into your mutant strain a DNA plasmid engineered to contain the DNA coding sequence of the human hexokinase gene, what must you also include? a)The human hexokinase promoter b)The bacterial hexokinase promoter c)Both the human and bacterial promoters d)You cannot engineer a bacteria to produce a human enzymearrow_forwardShown below is a portion of a wild-type DNA sequence that encodes the last amino acids of a protein that is 270 amino acids long. The first three bolded base pairs indicate the frame and include the coding region. 5^ ...GCTAAGTATTGCTCAAGATTAGGATGATAAATAACTGG 3^ 3^.. CGATTCATAACGAGTTCTAATCCTACTATTTATTGACC 5^ Which strand is the template strand for transcription of this gene? Briefly explain how you know. An insertion of one base pair causes the protein to decrease in length by seven amino acids. With respect to the sequence given above, where does this insertion occur? A change of one base pair leads to the protein increasing in the length by one amino acid. With respect to the sequence given above, which base pair would you change, and what would you change this base pair for the protein to increase in the length by one amino acid?arrow_forward
- You want to clone the Human Gene A into a plasmid for producing the Protein A in bacteria. GeneA encodes an mRNA of 100 nucleotides long, but the entire gene spans more than 4000 nucleotides. There are three exons and two introns. If we were to clone this gene directly from the nuclear DNA, bacteria would not be able to express the insulin protein. Explain why this is true.arrow_forwardConsider the following sequence of DNA: 3'-TTA CGG-5'What dipeptide is formed from this DNA after transcription and translation? b. If a mutation converts CGG to CGT in DNA, what dipeptide is formed? c. If a mutation converts CGG to CCG in DNA, what dipeptide is formed? d. If a mutation converts CGG to AGG in DNA, what dipeptide is formed?arrow_forwardConsider the Rho-dependent terminator sequence 5’CCCAGCCCGCCUAAUGAGCGGCCUUUUUUUU-3’. What affect would a point mutation at any one of the bolded and underlined nucleotides disrupt termination of transcription? Group of answer choices Mutation in one of these nucleotides would disrupt base pairing, preventing the formation of the hairpin and disrupting termination. Mutation in one of these nucleotides would have no affect on base pairing, so the termination hairpin is formed and termination proceeds. Mutation in one of these nucleotides would not disrupt base pairing, but would prevent the formation of the hairpin and disrupt termination. Mutation in one of these nucleotides would disrupt base pairing, but not affect the formation of the hairpin and termination proceeds.arrow_forward
- The following represent deoxyribonucleotide sequences in the template strand of DNA: Sequence 1: 5′-CTTTTTTGCCAT-3′ Sequence 2: 5′-ACATCAATAACT-3′ Sequence 3: 5′-TACAAGGGTTCT-3′ (a) For each strand, determine the mRNA sequence that would be derived from transcription. (b) Using Figure 12–7, determine the amino acid sequence that is encoded by these mRNAs. (c) For Sequence 1, what is the sequence of the partner DNA strand?arrow_forwardA temperature-sensitive mutation is one in which the defect is not presented functionally until the temperature is raised. In the case described below, the enzymes function normally in bacteria at 37 °C, but are non-functional at 40 °C. Predict the detailed molecular consequences of a loss of function in a temperature-sensitive mutant for each of the following enzymes: a) DNA gyrase, b) DNA polymerase III, c) DNA ligase, d) DNA polymerase I.arrow_forwardWhat is the production of RNA called and what is the enzyme that catalyzes the process?What are the similarities and differences between the transcription process and the repli-cation processes?Concerning their biological function what is the difference between DNA and RNA? Is there any situation in which DNA is made based on a RNA template? If there is,explain with an example how it occurs and state the enzyme involved?What is the difference between plasma membrane and cell wall?arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
