
Concept explainers
A
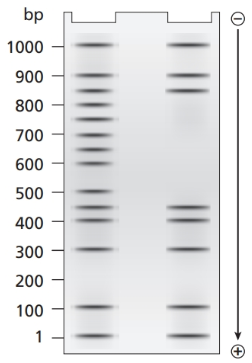
a. What length of DNA is bound by the transcriptional proteins? Explain how the gel results support this interpretation.
b. Draw a diagram of this DNA fragment bound by the transcriptional proteins, showing the approximate position of proteins along the fragment. Use the illustration style seen in Research Technique
c. Explain the role of DNase I.
Want to see the full answer?
Check out a sample textbook solution
Chapter 8 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- A linear piece of DNA that is 14 kb long is cut first by EcoRI alone, then by SmaI alone, and finally, by both EcoRI and SmaI together. The following results are obtained: Draw a map of the EcoRI and SmaI restriction sites on this 14-kb piece of DNA, indicating the relative positions of the restriction sites and the distances between them.arrow_forwardYou are engineering a new vector that contains a screenable marker that can be used for blue/white screening of successful clones. For each site (1, 2, and 3) on the cloning vector below, describe why it would or would not be a good place for you to put the polylinker to facilitate blue/white screening. You can assume that the polylinker itself will not interfere with coding sequence in that region. In other words, the polylinker length will be a multiple of 3 nucleotides, will not contain a stop codon, and any amino acids translated will not affect the activity of the protein in that region. The arrows indicate the direction of transcription for the gene.arrow_forwardYou are engineering a new vector that contains a screenable marker that can be used for blue/white screening of successful clones. For each site (1, 2, and 3) on the cloning vector below, describe why it would or would not be a good place for you to put the polylinker to facilitate blue/white screening. You can assume that the polylinker itself will not interfere with coding sequence in that region. In other words, the polylinker length will be a multiple of 3 nucleotides, will not contain a stop codon, and any amino acids translated will not affect the activity of the protein in that region. The arrows indicate the direction of transcription for the gene. Note from student:As stated in the problem... "YOU CAN ASSUME THAT THE POLYLINKER ITSELF WILL NOT INTERFERE WITH CODING SEQUENCE IN THAT REGION. IN OTHER WORDS, THE POLYLINKER LENGTH WILL BE A MULTIPLE OF 3 NUCLEOTIDES, WILL NOT CONTAIN A STOP CODON, AND ANY AMINO ACIDS TRANSLATED WILL NOT AFFECT THE ACTIVITY OF THE PROTEIN IN THAT…arrow_forward
- A lilP mutant called lilPXS is isolated that produces a truncated polypeptide of only 6 AA in length. Describe a single basepair DNA change that would lead to this truncated version of the protein. Multiple options are possiblearrow_forwardKnowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardAs part of a project investigating potential new drug targets in the fight against malaria, you are seeking to clone the gene for a protein from the malaria parasite Plasmodium falciparum. You wish to express this protein in BL21 (DE3) cells, a standard laboratory strain of Escherichia coli. After purification of your protein, you run an SDS-PAGE gel and notice that the major band has lower molecular weight than expected, so you fear you are getting a truncated version. 1. What technique could you use to confirm that you are obtaining a shortened version of your intended protein? explainarrow_forward
- As part of a project investigating potential new drug targets in the fight against malaria, you are seeking to clone the gene for a protein from the malaria parasite Plasmodium falciparum. You wish to express this protein in BL21 (DE3) cells, a standard laboratory strain of Escherichia coli. After purification of your protein, you run an SDS-PAGE gel and notice that the major band has lower molecular weight than expected, so you fear you are getting a truncated version. (a) Give TWO possible causes of your protein becoming truncated. explainarrow_forwardA linear piece of DNA that is 30 kb long is first cut with BamHI, then with HpaII, and, finally, with both BamHI and HpaII together. Fragments of the following sizes were obtained from this reaction: BamHI: 20-kb, 6-kb, and 4-kb fragments Hpall: 21-kb and 9-kb fragments BamHI and Hpall: 20-kb, 5-kb, 4-kb, and 1-kb fragments Draw a restriction map of the 30-kb piece of DNA, indicating the locations of the BamHI and HpaII restriction sitesarrow_forwardIn reversible terminator sequencing, how would the sequencing process be affected if the 3′-end-blocking group of each nucleotide were replaced with the 3′-H present in the dideoxynucleotides used in Sanger sequencing?arrow_forward
- You were going to sequence a rice DNA fragment whose sequence was only know at one end, as shown below. 5’ AAACGATCGAGTCGCATCCAAAATCGATACCC—unknown region 3’ TTTGCTAGCTCTGCGTAGGTTTTAGCTATGGG—unknown region After several tries, you obtained a beautiful sequencing image as shown here: The worked out well partially because you had designed a primer for sequencing the unknown region according to the following guideline: Tm is 55 – 60°C. Ensures primer had a appropriate melting temperature for PCR ans sequencing. The GC content of the primer is the same as the genome/template (rice = 60%, human/Drosophila = 45-50%). A same nucleotide cannot be more than 2 in a row, e.g. CCC, GGGGG, AAA. The secondary structure of the primer must be none or weak. No primer dimers (The primer anneals to itself). 3’ end is the most important: it should not end in A, preferably ends in GG, GC, CG or CC This website can help you design the primer: http://www.oligoevaluator.com/OligoCalcServlet…arrow_forwardIn the Avery, McLeod, McCarty Experiment where supernatant from heat killed, virulent S Strain pneumonia solutions were added to non-virulent R Strain pneumonia cell cultures and allowed to grow in liquid media (i.e., broth). In tubes where DNAase was added to the supernatant prior to cell culture, what was the observed effect when plating and growing the S. pneumonia cells to solid media? a All RNA was degraded and Transformation of the R Strain to S Strain occurred. b All Protein was degraded and Transformation of the R Strain to S Strain occurred. c All DNA was degraded and Transformation of the R Strain to S Strain did not occur. d All RNA was degraded and no Transformation occurred indicating RNA is the molecule of Transformation inheritance e None of the above are truearrow_forwardThat's the result of Gel electrophoresis of genomic DNA ( Of genomic DNA extraction experiment), please discuss the results and label and name the image to illustrate the answer? - Marker band sizes in gel: From top (well side) to bottom the bands have the following size in base-pair/bp- 6751,3652,2827,1568,1118,825,630arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

