
Genetics: From Genes To Genomes (6th International Edition)
6th Edition
ISBN: 9781260041217
Author: Leland Hartwell Dr., ? Michael L. Goldberg Professor Dr., ? Janice Fischer, ? Leroy Hood Dr.
Publisher: Mcgraw-Hill
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 9, Problem 15P
A recombinant DNA molecule is constructed using a plasmid vector called pMBG36 that is 4271 bp long. The pMBG36 plasmid contains a so-called polylinker that has a single site for each of several restriction enzymes, including BamHI (5′ G^GATCC 3′) and EcoRI (5′ G^AATTC 3′). The sequence of the polylinker region of pMBG36 is shown below; the dots indicate the large majority of the vector that is not shown. You now cut
pMBG36 with EcoRI and insert into it a fragment of the DNA previously shown in Problem 4 that is also cut with EcoRI.
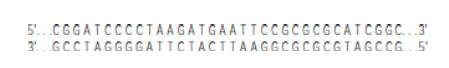
| a. | Write out as much of the DNA sequence of the resultant recombinant DNA molecule as is possible. Two answers are possible; you need to show only one. |
| b. | Why are there two possible answers to part (a)? |
| c. | How many recognition sites for BamHI will be found in the recombinant DNA molecule shown in your answer to part (a)? |
| d. | If you cut this recombinant DNA molecule with BamHI and run the digest on a gel, how many bands would you see and how large would they be? |
| e. | How many recognition sites for EcoRI will be found in the recombinant DNA molecule shown in your answer to part (a)? |
| f. | If you cut this recombinant DNA molecule with EcoRI and run the digest on a gel, how many bands would you see and how large would they be? |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Consider the following plasmid (size 8000 bp), with restriction sites at the positions indicated: (see image)
a) This plasmid is digested with the enzymes listed below. Indicate how many fragments will begenerated in each case, and give the sizes of the fragments.PstIXhoICombination of PstI + XhoI + EcoRI (triple digest)
b) Draw the banding pattern you would expect to observe if each of these digestions is loaded into a separate well of an agarose gel, and the fragments separated by electrophoresis. In the first well you load a DNA marker (M) containing fragments with sizes of 1000 bp, 2000 bp, 4000 bp and 8000 bp.
c) This gel is transferred to a membrane in a Southern blot experiment, and hybridised to a radioactively labelled 200 bp probe, which anneals to the plasmid DNA at the position indicated on the diagram above. Draw the autoradiographic profile you would expect to observe for the membrane.
A circular plasmid of 10,000 base pairs (bp) is digested with two restriction enzymes, A and B, to produce a
3000 bp and a 2000 bp bands when visualized on an agarose gel. When digested with one enzyme at a time,
only one band is visible at 5000 bp. If the first site for enzyme A (A1) is present at the 100h base, the order
in which the remaining sites (A2, B1 and B2) are present is -
(A) 3100, 5100, 8100
115.
(B) 8100, 3100, 5100
(C) 5100, 3100, 8100
(D) 8100, 5100,. 3100
A plasmid vector pBS281 is cleaved by the enzymeBamHI (5′ G^GATCC 3′), which recognizes only onesite in the DNA molecule. Human DNA is digestedwith the enzyme MboI (5′ ^GATC 3′), which recognizes many sites in human DNA. These two digestedDNAs are now ligated together. Consider only thosemolecules in which the pBS281 DNA has been joinedwith a fragment of human DNA. Answer the following questions concerning the junction between thetwo different kinds of DNA. a. What proportion of the junctions between pBS281and all possible human DNA fragments can becleaved with MboI?b. What proportion of the junctions between pBS281and all possible human DNA fragments can becleaved with BamHI?c. What proportion of the junctions between pBS281and all possible human DNA fragments can becleaved with XorII (5′ C^GATCG 3′)?d. What proportion of the junctions between pBS281and all possible human DNA fragments can becleaved with BstYI (5′ R^GATCY 3′)? (R and Ystand for purine and pyrimidine, respectively.)e.…
Chapter 9 Solutions
Genetics: From Genes To Genomes (6th International Edition)
Ch. 9 - Match each of the terms in the left column to the...Ch. 9 - For each of the restriction enzymes listed below:...Ch. 9 - The calculations of the average restriction...Ch. 9 - The DNA molecule whose entire sequence follows is...Ch. 9 - Why do longer DNA molecules move more slowly than...Ch. 9 - Agarose gels with different average pore sizes are...Ch. 9 - The following picture shows the ethidium...Ch. 9 - The linear bacteriophage genomic DNA has at each...Ch. 9 - Consider a partial restriction digestion, in which...Ch. 9 - The text stated that molecular biologists have...
Ch. 9 - a. What is the purpose of molecular cloning? b....Ch. 9 - a. DNA polymerase b. RNA polymerase c. A...Ch. 9 - Is it possible that two different restriction...Ch. 9 - A plasmid vector pBS281 is cleaved by the enzyme...Ch. 9 - A recombinant DNA molecule is constructed using a...Ch. 9 - Suppose you are using a plasmid cloning vector...Ch. 9 - Prob. 17PCh. 9 - The lacZ gene from E. coli encodes the enzyme...Ch. 9 - Your undergraduate research advisor has assigned...Ch. 9 - Which of the enzymes from the following list would...Ch. 9 - You use the primer 5 GCCTCGAATCGGGTACC 3 to...Ch. 9 - a. To make a genomic library useful for sequencing...Ch. 9 - Problem 15 showed part of the sequence of the...Ch. 9 - Eukaryotic genomes are replete with repetitive...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A closed circular plasmid B-DNA (10.5 bp/turn) consists of 231 base pairs and has Wr= -1 (ccDNAa). Then a topoisomerace acts upon ccDNAa leading to strain relaxation caused by supercoiling, thus, a topoisomere ccDNAb is formed. Finally, EtBr (Ethidium bromide) intercalator is added leading to ccDNAc which has 11bp/turn. a) Calculate superhelical densities σa, σb, σc of the three plasmids ccDNAa, ccDNAb and ccDNAc, respectively. b) Which of the three topoisomeres will move faster in agarose gel electrophoresis and why?arrow_forwardUsing the plasmid map of pBCH2.0 provided above, predict how many DNA fragments would be formed if this plasmid was digested with restriction enzyme BamHI.arrow_forwardThe restriction endonuclease NotI recognizes the octanucleotide sequence GCGGCCGC. Calculate the expected number of NotI cleavage sites in the bacteriophage λgenome, a linear DNA duplex 48.5 kbp in length with a (G + C) content of 50%.arrow_forward
- The partial sequence of one strand of a double-stranded DNA molecule is5′ – – – GACGAAGTGCTGCAGAAAGTCCGCGTTATAGGCATGAATTCCTGAGG – – – 3′The cleavage sites for the restriction enzymes EcoRI and PstI are shown below.Write the sequence of both strands of the DNA fragment created when this DNA is cleaved with both EcoRI and PstI. The top strand of your duplex DNA fragment should be derived from the strand sequence given abovearrow_forwardKpn I and Acc 65I are restriction enzymes that identify and cleave the same 6-bp sequence. The sticky end created by Kpn I cleavage, on the other hand, cannot be directly ligated to the sticky end formed by Acc 65I cleavage. Please explain why.arrow_forwardGiven the following double-stranded fragment of DNA: 5'- ACTTGGCAGGCCTTCGATCC-3' 3'- TGAАССGTCСGGAAGCTAGG-5' A hypothetical restriction endonuclease recognizes a 6bp sequence with two-fold symmetry (typical for restriction enzymes) found in this fragment and catalyzes cleavage of this DNA on both strands between GG nucleotides within the recognition sequence. This nuclease exhibits b-type cleavage (atypical for restriction enzymes). Draw the double-stranded sequence of each fragment after cleavage showing any phosphates left on the ends.arrow_forward
- The restriction endonuclease NciI recognizes and cuts the five-base-pair sequence 5’- CC(G/C)GG-3’ [where (G/C) means either G or C will work at that position]. (1) How often, on average, would this sequence occur in random DNA? Assume the DNA contains 25% each of A, G, T & C. (2) After digestion, Nci1 leaves a one-base 5’ overhang. Write/draw the cut site/digested products.arrow_forwardThe restriction enzymes Kpn I and Acc 65I recognize and cleave the same 6-bp sequence. However, the sticky end formed from Kpn I cleavage cannot be ligated directly to the sticky end formed from Acc 65I cleavage. Explain why.arrow_forwardExamine the structure of the pBR322 plasmid depicted below. Assume total size of the plasmid is 4,361 bp and the blue numbers indicate locations of restriction sites relative to the O point at the top of the plasmid. What size fragments would be generated by the following restriction digestion reactions? 1. Sall 2. Sal 1 + BamH1 3. Sal 1 + EcoR1 4. Sal I + BamH1 + EcoR1 PstI 3607 3000 4000 amp ori HindIII Edit View Insert Format Tools Table EcoRI EcoRV 4359 0 29 185 pBR322 4361 bp 2295 NdeI tet 2000 BamHI 375 651 SalI 1000 Type a short answer in the space provided below.arrow_forward
- a)Dr. Thisisaneasyexam decides to amplify a gene from a plasmid using PCR. She starts out with 6.6 x 10-14g of a 10 kb template in a 100 µl reaction. Assuming that the molecular weight of the average base pair is 660 Daltons calculate the number of molecules of the template. b)Given that the concentration of each primer (20 base pairs each) is 0.1µM in this same reaction volume (100 µl) calculate the number of molecules of the primer presentarrow_forwardThe partial sequence of one strand of a double-stranded DNA molecule is 5'-GACGAAGTGCTGCAGAAAGTCCGCGTTATAGGCATGAATTCCTGAGG -3' EcoRI is a restriction enzyme that cleaves after G in the sequence 5'-GAATTC-3'. PstI is a restriction enzyme that cleaves after A in the sequence 5'-CTGCAG-3'. Write the sequence of both strands of the DNA fragment created when this DNA is cleaved with both EcoRI and PstI. The first strand of your duplex DNA fragment should be derived from the given strand sequence. 5'- -3' 3'- -5'arrow_forwardYou are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), Hindlll (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. +18 a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license