
Concept explainers
Cartoon renderings of the proteins Top 7 and adaH2 are shown below. Both are soluble, densely packed proteins of roughly 96 residues, and each has a topology of 2 a helices packed onto a 5-stranded ß sheet.
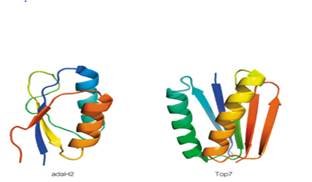
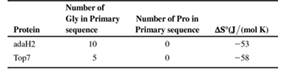
The accompanying table lists some information about the amino acid composition and values of ΔS0 for the folding of these proteins (i.e. for Unfolded →Folded) at 250C. Based on the information in the table, which protein do you predict buries the greater hydrophobic surface area upon folding? Assume 2-state folding (i.e., no intermediates), and that the Unfolded state for both proteins is 100% solvent-exposed. Explain your answer in terms of expected contributions from ΔS0peptide and ΔS0solvent.
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
- You are given a protein solution with a concentration of 0.15 mg/ml. Suppose that we want to prepare a solution containing 100 μg of the protein at a concentration of 1 mg/ml. To achieve this, we will first dry down enough protein solution to obtain 100 μg of proteins. How much solution do we need for drying down? How much volume of H2O do we need to add to the dried protein to obtain the desired concentration?arrow_forwardWhich intermolecular forces are important in acetic acid, CH3 –(C=0)-oh? A particular amino acid contains a- CHNH3+ group. Is this amino acid more likely to be found on the inside or the outside of the folded protein? Briefly explain. The addition of ethanol, CH3CHOH, t an aqueous solution lowers the surface tension of the solution. Predict whether adding ethanol to an aqueous protein solution will tend to stabilize or unfold the protein. Briefly explain.arrow_forward. Disulfide bonds have been shown to stabilize proteins (i.e., make them less likely to unfold). Consider the cases shown schematically below for two variants of the same protein. In case #1 the disulfide forms between Cys residues that have been introduced near the protein N- and C-termini, and in case #2 the disulfide forms between Cys residues that have been introduced in the middle of the protein sequence. Which protein is likely to be more stable? (Note: Assume the disulfide bond is intact in both the unfolded and folded states). Explain your reasoning.arrow_forward
- The following are sequences from three different alpha helices found inhuman proteins: hBak(72-87): GQVGRQLAIIGDDINR hCB1(196-210): VTASFTASVGSLFLT hCB2(248-262): LVLAVLLICWFPVL Classify each of these three helices as either a) mostly hydrophobic, b)mostly hydrophilic, or c) amphipathic. Use the helical wheel to explainyour answers. Given the character of these helices, in which part of the protein wouldyou expect them to reside?arrow_forwardhow does the protein environment surrounding an amino acid chain affect its chemical properties?Consider the carboxyl group on an asparate side chain in the following environments in a protein. Rank in order these environments from the highest to the lowest proportion of carboxyl groups in the -COO- form.that is in terms of pKas. 1. an aspartate side chain on the surface of protein with no other ionizable groups nearby. 2. an aspartate side chain buried in a hydrophobic pocket on the surface of a protein 3. an asparate chain in a hydrophobic pocket adjacent to a glutamte side chain 4. an asparate side chain in a hydrophobic porket adjacent to a lysine side chain.arrow_forwardBoth hsp60-like and hsp70 molecular chaperonesshare an affinity for exposed hydrophobic patches on pro-teins, using them as indicators of incomplete folding. Whydo you suppose hydrophobic patches serve as critical sig-nals for the folding status of a protein?arrow_forward
- Do you expect a Pro → Gly mutation in a surface-loop region of a globular protein to be stabilizing or destabilizing? Assume the mutant folds toa native-like conformation. Explain your answer in terms of the predictedenthalpic and entropic effects of the mutation on the ∆G for protein foldingcompared to ∆G of folding for the wild-type proteinarrow_forwardConsidering the chemical characteristics of the amino acids valine and glutamic acid (see Figure 5.14), propose a possible explanation for the dramatic effect on protein function that occurs when valine is substituted for glutamic acid.arrow_forwardCurrently, aspartic acid is forming an ionic interaction with arginine in a protein. Part a) If arginine is replaced with glutamic acid, would the ionic interaction have its stability increased, decreased, or have no effect on the ionic interaction? Part b) If arginine is replaced with Lysine, would the ionic interaction have its stability increased, decreased, or have no effect on the ionic interaction? Part c) If arginine is replaced with isoleucine, would the ionic interaction have its stability increased, decreased, or have no effect on the ionic interaction?arrow_forward
- The major difference between a protein molecule in its native state and inits denatured state lies in the number of conformations available. To a firstapproximation, the native, folded state can be thought to have one conformation. The unfolded state can be estimated to have three possible orientations about each bond between residues.(a) For a protein of 100 residues, estimate the entropy change per moleupon denaturation.(b) What must be the enthalpy change accompanying denaturation to allow the protein to be half-denatured at 50 °C?(c) Will the fraction denatured increase or decrease with increasingtemperature?arrow_forwardRemembering that the amino acid side chains projecting from each polypeptide backbone in a β sheet point alternately above and below the plane of the sheet, consider the following protein sequence: Leu-Lys-Val-Asp-Ile-Ser-Leu-Arg- Leu-Lys-Ile-Arg-Phe-Glu. Do you find anything remarkable about the arrangement of the amino acids in this sequence when incorporated into a β sheet? Can you make any predictions as to how the β sheet might be arranged in a protein?arrow_forwarda) Canonical forces in protein folding. Describe how these forces come into play when a protein folds. Why do are other intermolecular interactions important to fully understand folding processes?arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





