
Concept explainers
The phage P1 is used as a generalized transducing phage in an experiment combining a donor strain of E. coli of genotype leu+phe+ala+ and a recipient strain that is leu-phe-ala-. In separate experiments, transductants are selected for leu+ (experiment A), for phe+ (experiment B), and for ala+ (experiment C). Following selection, transductant genotypes for the unselected markers are identified.
a. What compound or compounds are added to the minimal medium to select for transductants in experiments A, B, and C?
b. Selection experiment results below show the frequency of each genotype.
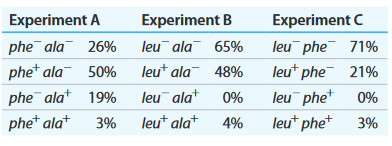
c. Determine the order of genes on the donor chromosome.
d. Diagram the crossover events that form each of the transductants in experiment A.
e. In experiment B, why are there no transductants with the genotype leu- ala+?
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Study Guide And Solutions Manual For Genetic Analysis: An Integrated Approach
- What is meant by the term site-specific recombination as used in identifying the processes that lead to the integration of temperate bacteriophages into host bacterial chromosomes during lysogeny or to the formation of specialized transducing phage?arrow_forwardIn Hershey-Chase experiment, bacteriophages protein coats were tagged with radioactive isotope S-32. These phages were used to infect E. coli cells and the cells were further centrifuged to form pellets. Why was the radioactivity level of S-32 found greater outside the cells compared to the E. coli cell pellets? Explain briefly. If the experiment is repeated in the same manner but this time the phage protein coats are labelled with isotope X and the phage DNA with isotope Y, which isotope’s radioactivity will be found in greater amounts in the E. coli cell pellets after centrifugation? Explain briefly.arrow_forwardIn a generalized-transduction experiment, phages arecollected from an E. coli donor strain of genotype cys+leu+ thr+ and used to transduce a recipient of genotypecys- leu- thr-. Initially, the treated recipient populationis plated on a minimal medium supplemented with leucine and threonine. Many colonies are obtained.a. What are the possible genotypes of these colonies?b. These colonies are then replica plated onto threedifferent media: (1) minimal plus threonine only, (2)minimal plus leucine only, and (3) minimal. Whatgenotypes could, in theory, grow on these three media?c. Of the original colonies, 56 percent are observed togrow on medium 1, 5 percent on medium 2, and nocolonies on medium 3. What are the actual genotypes ofthe colonies on media 1, 2, and 3?d. Draw a map showing the order of the three genes andwhich of the two outer genes is closer to the middle genearrow_forward
- In five Hfr strains, each of which was used to build a time-of-transfer map, the genes entered the recipient cells as follows: Strain 1: S L A C T F Strain 2: N P F T C A Strain 3: T F P N U Y Strain 4: S H Y U N P Strain 5: U N P F T C Which of the following represents a correct gene map of these results? N P F T S L A C H U Y S L A C T F P N H Y U C T F P N U Y H S L A T C A L S P N U Y H F U N P C A L S F T H Yarrow_forwardDoes the Hershey and Chase experiment rule out the possibility that RNA is the genetic material of T2 phage? Explain. If it does not, redesign the experiments of Hershey and Chase to distinguish between DNA and RNA in the T2 phage.arrow_forwardThe bacteriophage genome consists of many genes encoding proteins that make up the head, collar, tail, and tail fibers. When these genes are transcribed following phage infection, how are these proteins synthesized, since the phage genome lacks genes essential to ribosome structure?arrow_forward
- In E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What information do you know based onthe question and your understanding of the topic?arrow_forwardIn E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What topic in genetics does this question address?arrow_forwardBy conducting conjugation experiments between Hfr and recipientstrains, Wollman and Jacob mapped the order of many bacterialgenes. Throughout the course of their studies, they identified severaldifferent Hfr strains in which the F-factor DNA had been integratedat different places along the bacterial chromosome. A sample of theirexperimental results is shown in the table:Draw a map that shows the order of genes and the locations ofthe origins of transfer among these different Hfr strains?arrow_forward
- Two mutations that affect plaque morphology in phages (a− and b −) have been isolated. Phages carrying both mutations (a− b−) are mixed with wild-type phages (a+ b+) and added to a culture of bacterial cells. Once the phages have infected and lysed the bacteria, samples of the phage lysate are collected and cultured on plated bacteria. The following numbers of plaques are observed: Plaque phenotype Number a+ b+ 2043 a+ b− 320 a− b+ 357 a− b− 2134 What is the frequency of recombination between the a and b genes?arrow_forwardA researcher is studying the rII locus of phage T4. FourrII− strains are obtained: A, B, C, and D. In the first experiment, E. coli strain K(λ) is coinfected with two rII− strains simultaneously and the results are recorded. Infection with A and B phage = lysis occurs Infection with A and C phage = lysis occurs Infection with B and C phage = no lysis occurs Infection with B and D phage = no lysis occurs Infection with C and D phage = no lysis occurs In a second experiment, coinfections are performed first in E. coli strain B, then the progeny phage are used to infect E. coli strain K(λ). Progeny of A and B phage = plaques form Progeny of B and C phage = plaques form Progeny of C and D phage = plaques form Progeny of B and D phage = no plaques from Which conclusions are consistent with these data? Why? A) Strains A and B carry mutations in the same gene. B) Strains B and D both carry the same mutation. C) Strains B, C, and D carry mutations in the same gene. D)…arrow_forwardThe bacteriophage genome consists primarily of genes encodingproteins that make up the head, collar and tail, and tail fibers When these genes are transcribed following phage infection, howare these proteins synthesized, since the phage genome lacksgenes essential to ribosome structure?arrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
