
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 8, Problem 5P
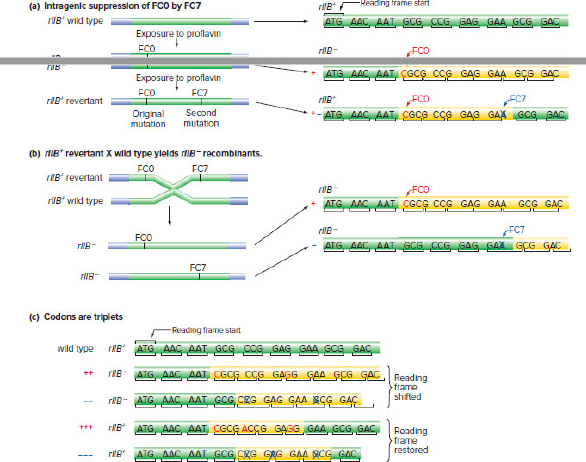
Consider Crick and Brenner’s experiments in Fig. 8.4, which showed that the genetic code is based on
| a. | Crick and Brenner obtained FC7, an intragenic suppressor of FC0, that was a mutation in a second site in the rIIB gene near the FC0 mutation. Describe a different kind of mutation in the rIIB gene these researchers might have recovered by treating the FC0 mutant with proflavin and looking for restored rIIB+ function. |
| b. | How could Crick and Brenner tell the difference between the occurrence described in part (a) and an intragenic suppressor like FC7? |
| c. | When FC7 was separated from FC0 by recombination, the result was two rIIB mutant phages: One was FC7 and the other was FC0. How could they discriminate between the rIIB recombinants that were FC7 and those that were FC0? |
| d. | Explain how Crick and Brenner could obtain different deletion (−) or addition (+) mutations so as to make the various combinations such as ++, −−, +++, and −−− shown in Fig. 8.4c. |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Ochre and amber are two distinct nonsense mutations. Before the genetic code was worked out, Sydney Brenner, Anthony O. Stretton, and Samuel Kaplan applied different types of mutagens to bacteriophages in an attempt to determine the bases present in the codons responsible for amber and ochre mutations. They knew that the ochre and amber mutations were suppressed by different types of suppressor mutations, which demonstrated that each is a different stop codon. They obtained the following results: (1) A single-base substitution could convert an ochre mutation into an amber mutation. (2) Hydroxylamine induced both ochre and amber mutations in wildtype phages. (3) 2-Aminopurine caused ochre to mutate to amber. (4) Hydroxylamine did not cause ochre to mutate to amber. These data do not allow the complete nucleotide sequence of the amber and ochre codons to be worked out, but they do provide some information about the bases found in the nonsense mutations. a. What conclusions about the…
Ochre and amber are two distinct nonsense mutations. Before the genetic code was worked out, Sydney Brenner, Anthony O. Stretton, and Samuel Kaplan applied different types of mutagens to bacteriophages in an attempt to determine the bases present in the codons responsible for amber and ochre mutations. They knew that the ochre and amber mutations were suppressed by different types of suppressor mutations, which demonstrated that each is a different stop codon. They obtained the following results: (1) A single-base substitution could convert an ochre mutation into an amber mutation. (2) Hydroxylamine induced both ochre and amber mutations in wildtype phages. (3) 2-Aminopurine caused ochre to mutate to amber. (4) Hydroxylamine did not cause ochre to mutate to amber. These data do not allow the complete nucleotide sequence of the amber and ochre codons to be worked out, but they do provide some information about the bases found in the nonsense mutations.
Q. Of the three nonsense codons…
A mutant strain of Salmonella bacteria carries a mutation of the rho protein that has fully activity at 37°C but is completely inactivated when the mutant strain is grown at 40°C.
a)Speculate about the kind of differences you would expect to see if you compared a broad spectrum of mRNAs from the mutant strain grown at 37°C and the same spectrum of mRNAs from the strain when grown at 40°C.
b)Are all the mRNAs affected by the rho protein mutation in the same way? Why or why not?
Chapter 8 Solutions
Genetics: From Genes to Genomes
Ch. 8 - For each of the terms in the left column, choose...Ch. 8 - Match the hypothesis from the left column to the...Ch. 8 - How would the artificial mRNA 5GUGUGUGU . . . 3 be...Ch. 8 - An example of a portion of the T4 rIIB gene in...Ch. 8 - Consider Crick and Brenners experiments in Fig....Ch. 8 - The HbSsickle-cell allele of the human -globin...Ch. 8 - The following diagram describes the mRNA sequence...Ch. 8 - The amino acid sequence of part of a protein has...Ch. 8 - The results shown in Fig. 8.5 may have struck you...Ch. 8 - Identify all the amino acid-specifying codons in...
Ch. 8 - Before the technology existed to synthesize RNA...Ch. 8 - A particular protein has the amino acid sequence...Ch. 8 - How many possible open reading frames frames...Ch. 8 - Prob. 14PCh. 8 - Charles Yanofsky isolated many different trpA-...Ch. 8 - The sequence of a segment of mRNA, beginning with...Ch. 8 - You identify a proflavin-generated allele of a...Ch. 8 - Using recombinant DNA techniques which will be...Ch. 8 - Describe the steps in transcription that require...Ch. 8 - Chapters 6 and 7 explained that mistakes made by...Ch. 8 - The coding sequence for gene F is read from left...Ch. 8 - If you mixed the mRNA of a human gene with the...Ch. 8 - Prob. 23PCh. 8 - The Drosophila gene Dscam1 encodes proteins on the...Ch. 8 - Describe the steps in translation that require...Ch. 8 - Locate as accurately as possible the listed items...Ch. 8 - Concerning the figure for Problem 26: a. Which...Ch. 8 - a. Can a tRNA exist that has the anticodon...Ch. 8 - For parts a and b of Problem 28, consider the DNA...Ch. 8 - Remembering that the wobble base of the tRNA is...Ch. 8 - Prob. 31PCh. 8 - The yeast gene encoding a protein found in the...Ch. 8 - The sequence of a complete eukaryotic gene...Ch. 8 - Arrange the following list of eukaryotic gene...Ch. 8 - Prob. 35PCh. 8 - The human gene for 2 lens crystallin has the...Ch. 8 - In prokaryotes, a search for genes in a DNA...Ch. 8 - a. The genetic code table shown in Fig. 8.2...Ch. 8 - a. Very few if any eukaryotic genes contain tracts...Ch. 8 - Explain how differences in the initiation of...Ch. 8 - Do you think each of the following types of...Ch. 8 - Null mutations are valuable genetic resources...Ch. 8 - The following is a list of mutations that have...Ch. 8 - Considering further the mutations described in...Ch. 8 - Adermatoglyphia described previously in Problem 18...Ch. 8 - Prob. 46PCh. 8 - You learned in Problem 21 in Chapter 7 that the...Ch. 8 - When 1 million cells of a culture of haploid yeast...Ch. 8 - Why is a nonsense suppressor tRNATyr, even though...Ch. 8 - A mutant B. adonis bacterium has a nonsense...Ch. 8 - You are studying mutations in a bacterial gene...Ch. 8 - Another class of suppressor mutations, not...Ch. 8 - Yet another class of suppressor mutations not...Ch. 8 - At least one nonsense suppressing tRNA is known...Ch. 8 - An investigator was interested in studying UAG...Ch. 8 - Prob. 56PCh. 8 - In certain bacterial species, pyrrolysine Pyl,...Ch. 8 - Canavanine is an amino acid similar to arginine...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- a. Some antibiotics, such as rifampin, interfere with the function of RNA polymerase. What biological process is rifampin disrupting? b. Some antibiotic-resistant M. tuberculosis bacteria have a single point mutation (CàT) in the rpoB gene that causes an amino acid change from serine (a polar amino acid) to leucine (a non-polar amino acid). What type of mutation is this? Do you expect this to have no effect, a small effect, or a large effect on the polypeptide produced? Explain your reasoning. c. The rpoB gene encodes a subunit of the bacterial RNA polymerase protein. The point mutation described in Question 2 causes a change in protein folding, which leads to the inability of the rifampin antibiotic to bind to the RNA polymerase. Which level(s) of protein structure is/are affected by this change?arrow_forwardBased on the following wild type sequence, indicate if each of the mutations should be classified as : insertion, deletion, missense, nonsense, silent (Use the provided Genetic Code table). Wild Type: GUC GCC GAC GAG AGG Mutant 1: GUC GCC AGA CGA GAG G Mutant 2: GUC GCC ACG AGA GG Mutant 3: GUC GCC AAC GAG AGG Mutant 4: GUA GCC GAC GAG AGG Mutant 5: GUC GCC GAC UAG AGGarrow_forward. Let’s say that you have incredible skill and can isolate the white and red patches of tissue from the Drosophila eyes shown in Figure 12-24 in order to isolate mRNA from each tissue preparation. Using your knowledge of DNA techniques from Chapter 10, design an experiment that would allow you to determine whether RNA is transcribed from the white gene in the red tissue or the whitetissue or both. If you need it, you have access to radioactive white-gene DNAarrow_forward
- A graduate student studying the pathogenic bacteria Acinetobacter baumannii made cDNA from planktonic cells and biofilm cells and performed RNA-Seq on both samples. She aligned her sequencing reads to a locus of the baumannii genome as shown. a. Which genes are on an operon together? Explain which data supports this? b. What is the most expressed transcript from the locus in Planktonic culture? c. Order the transcripts from largest increase in relative expression between biofilm and planktonic cells to the largest decrease in relative expression. d. When this data was compared to microarray transcriptional profiling, the microarray data provided lower expression levels for the most highly expressed transcripts. Why did this occur?arrow_forwardGeneticists often use ethylmethane sulfonate (EMS) to induce mutations in Drosophila. Why is EMS a mutagen of choice for genetic research? What would be the effects of EMS in a strain of Drosophila lacking functional mismatch repair systems?arrow_forwardThe genetic alteration responsible for sickle-cell anemia in humans involves: a transition mutation from A to G, substituting glutamic acid for valine in a-globin a transversion mutation from T to A, substituting valine for glutamic acid in b-globin a transition mutation from T to C, substituting valine for glutamic acid in b-globin a transversion mutation from G to C, substituting glutamic acid for valine in a-globin a frameshift mutation of one ATC codon, removing glutamic acid from b-globinarrow_forward
- As we described in class, in the early 1960's Francis Crick and colleagues set out to determine how many nucleotide bases make up a codon, before it was possible to sequence DNA and before Nirenberg and his colleagues solved the genetic code. To do this, they used a chemical mutagen that they knew made single nucleotide changes, used this mutagen to conduct a screen for mutations that disrupted a particular gene, and collected a number of different mutations in this gene. Briefly describe the logic they used to deduce that the codon length is 3 nucleotides long.arrow_forward. Another class of suppressor mutations, not describedin the chapter, are mutations that suppress missensemutations.a. Why would bacterial strains carrying such missense suppressor mutations generally grow moreslowly than strains carrying nonsense suppressormutations?b. What other kinds of mutations can you imagine ingenes encoding components needed for gene expression that would suppress a missense mutationin a protein-coding gene?arrow_forwardHow long would it take for the E. coli RNA polymerase to synthesize the primary transcript for the E. coli genes encoding the enzymes for lactose metabolism, the 5,300 bp5,300 bp lac operon? Assume an average elongation rate of 7070 nucleotides per second. a)How far along the DNA would the transcription "bubble" formed by RNA polymerase move in 10 seconds10 seconds? b)Assuming that human Pol II transcribes at a similar rate, how long does it take to transcribe the 2,000,000 bp2,000,000 bp dystrophin gene?arrow_forward
- Albinism is a condition where individuals can't make melanin pigment. The affected gene encodes for tyrosine aminotransferase, a key enzyme in melanin production. If you analyze the DNA of an albino individual, of the following mutations, which one is the least likely mutation responsible for the albino phenotype? A. A substitution of A to G at 3' splicing site. B. A deletion in the TATA box region. C. An insertion after the start codon. D. An extra stretch of TTAATT in intron 1.arrow_forwardPTC is not a biological compound, but a chemical which became known after research done by Arthur J. Fox. Why is the TAS2R38 gene also called the PTC gene?arrow_forwardConsider the Rho-dependent terminator sequence 5’CCCAGCCCGCCUAAUGAGCGGCCUUUUUUUU-3’. What affect would a point mutation at any one of the bolded and underlined nucleotides disrupt termination of transcription? Group of answer choices Mutation in one of these nucleotides would disrupt base pairing, preventing the formation of the hairpin and disrupting termination. Mutation in one of these nucleotides would have no affect on base pairing, so the termination hairpin is formed and termination proceeds. Mutation in one of these nucleotides would not disrupt base pairing, but would prevent the formation of the hairpin and disrupt termination. Mutation in one of these nucleotides would disrupt base pairing, but not affect the formation of the hairpin and termination proceeds.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Mitochondrial mutations; Author: Useful Genetics;https://www.youtube.com/watch?v=GvgXe-3RJeU;License: CC-BY