
Concept explainers
The map below illustrates three alleles in a genome segment. The alleles differ in the number and location of restriction sequences. Restriction digestion and Southern blotting with the molecular probe whose hybridization location is indicated results in detection of a single DNA
band for each allele.
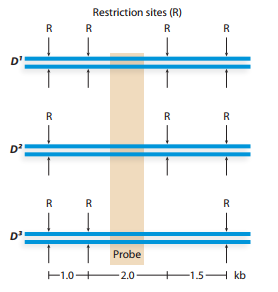
List the size (in kilobases) of DNA bands detected by Southern blotting of restriction digested DNA from organisms with the genotypes
Restriction
Organisms with the genotype
Why does this genotype produce a single detectable band, and why are the
Want to see the full answer?
Check out a sample textbook solution
Chapter 10 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- DNA samples from four individuals were cleaved with the same MW restriction endonuclease. The DNA fragments were separated by gel clectrophoresis, transferred to a membrane, and hybridized with a 12 kb 10 kb DNA probe complementary to a region between sites C and D (see 8 kb hybridization line). The image of the southern blot shows the labeled DNA bands and 6 kb molecular weight (MW) markers. The lane labels I, II, III, and IV -5 kb correspond to individuals I, II, III, and IV. Assume that fragments such as C-D and C-E are clearly resolved in this gel system. Fragment sizes are as given: A-B is 4 kb, B-C is I kb, C-D is 5 kb, and D-E is 650 bp. Individual I has five cleavage sites (A, B, C, D, and E) for the restriction endonuclease. DNA homologous to probe Which individual has at least one point mutation that eliminates restriction site C only? O II IV cannot be determined II Which individual has at least one point mutation that climinates restriction sites B and C? III O IV cannot be…arrow_forwardSickle cell anaemia is caused by a mutation in the HBB gene. The normal wild-type βA allele contains an MstII restriction site; in the mutated sickle-cell βS allele this restriction site has been lost.PCR amplification of part of the gene encompassing the mutation site from a normal unaffected individual results in a product of 110bp. Digestion of the PCR product with MstII and subsequent gel electrophoresis results in a single band of 55bp. How many bands and of what size would you expect to see in individuals who: i) has sickle-cell anaemia (has two βS alleles) ii) carry the disease (sickle cell trait) (has one βS and one βA allele) a. Individual Number of bands Size of bands (bp) Sickle-cell anaemia(has two βS alleles) 2 55 Sickle cell trait)(has one βS and one βA allele) 1 110 b. Individual Number of bands Size of bands (bp) Sickle-cell anaemia(has two βS alleles) 2 110 Sickle cell trait)(has one βS and one βA allele) 2 55 & 110…arrow_forwardAll of the following are performed during restriction fragment length polymorphism analysis. 1. splitting of double-stranded into single-stranded DNA 2. gel electrophoresis 3. autoradiography 4. immersion in radioactive probes 5. digestion of DNA with restriction endonucleases 6. use of a positive charge to transfer single-stranded DNA from a gel to a membrane. The correct sequence of these operations is whatarrow_forward
- The following DNA sequence has been identified and named ‘Gene Z’. It is thought that this sequence codes for a single protein. Design 5’ and 3’ PCR primers that will amplify ONLY the open reading frame of the genes. Give these sequences in a 5’ to 3’ orientation. Include EcoR1 and NotI restriction enzyme recognition sites on the 5’ ends of the 5’ and 3’ primers respectively.arrow_forwardThe map of plasmid pUC19 is shown below. The restriction site coordinate is the position of the 5’base on the top strand of each site sequence. The restriction enzyme sites are in bold type if there is only one site in pUC19. Please list the fragments in order of size, largest to smallest, which will result from a complete digestion by the restriction enzyme PvuII. Please list the fragments in order of size, largest to smallest, which will result from a complete digestion by the restriction enzyme DrdI.arrow_forwardIn the following "gene library" cloning experiment Digested genomic DNA AmpR gene TCR gene TCR is tetracycline resistant marker, AmpR is ampicillin resistant marker and BamHI is the unique restriction enzyme on plasmid. A PhD student digests/cuts the plasmids with BamHI restriction enzyme and the genomic DNA with EcoRI restriction enzyme. After performing the cloning experiment and obtaining colonies on a selection plate, the obtained cells will be ..... (Hint: this question is even more challenging; the PhD student was later demoted to an MSc student). a) resistant to ampicillin and tetracycline b) sensitive to tetracycline and ampicillin c) resistant to tetracycline and sensitive to ampicillin d) resistant to ampicillin and sensitive to tetracycline e) sensitive to ampicillin and tetracycline BamHIarrow_forward
- A group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardWhich RE can be used to distinguish bacterial strain 1 from strain 2 based on a sample DNA fragment from each strain (as shown below)? STRAIN 1 ~712 bp STRAIN 2 ~712 bp Restriction enzymes: O BglIII, Ndel, and Ncol O Ncol O Ndel OBgIIII CATATGCCATGGAGATCT GTATACGGTACCTCTAGA ~231 bp CACATGCCATGGAGATCT GTGTACGGTACCTCTAGA 231 bp BglIII Ncol AGATCT CCATGG TCTAGA GGTACC Nde1 CATATG GTATACarrow_forwardAn individual is heterozygous for an allele. Allele A has a SNP that results in disruption of an Not1 restriction enzyme site. Allele B has a functional Not1 restriction enzyme site. PCR is performed on the region of the gene containing the SNP to produce a 255bp PCR product. Restriction digestion was subsequently performed using EcoR1 restriction enzyme and the fragments were electrophoresed on an agarose gel. (EcoR1 cuts allele B at a site 115bp from the end of the sequence.) What are the sizes of the subsequent DNA fragments? 1) 115bp + 140bp 2) 205bp + 255bp 3) 140bp 4) 205bp + 255bp + 115bp 5) 140bp + 205bp + 255bp 6) 255bp + 115bp + 140bp 7) 115bp 8) 255bp 9) 115bp + 255bp 10) 255bp + 115bp + 140bp + 205bp 11) 140bp + 205bp 12) 205bp 13) 205bp + 115bp 14) 140bp + 255bparrow_forward
- The right-hand column contains descriptions of 5 types of molecular markers. Label each description with the name of the being described. Single-nucleotide polymorphism (SNP) Sequence-tagged site (STS) Microsatellite Amplified restriction fragment length polymorphism (AFLP) Restriction fragment length polymorphism (RFLP) Reset Zoom TABLE 23.1 Common Types of Molecular Markers Marker Description A site in a genome where the distance between two restriction sites varies among different individuals. These sites are identified by restriction enzyme digestion of chromosomal DNA and the use of Southern blotting. Amplified restriction fragment length polymorphism (AFLP) The same as an RFLP except that the site is amplified via PCR instead of isolating the chromosomal DNA. A site in the genome that contains many short sequences that are repeated many times in a row. The total length is usually in the range of 50-200 bp, and the length of a given microsatellite may be polymorphic within a…arrow_forwardConsider the following image of a stained agarose gel representing the separation of a plece of DNA that has been digested by a series of restriction enzymes. Assume no digest fragment is of identical size to another digest fragment. nucleotide pairs (kb) 8 S 45 3.5 AA size Hindill + markers EcoRI Hindill EcoRI - 1st attempt Based on all data presented in the stained agarose gel, which of the following statements is correct? Choose one: OA. The original piece of DNA was circular and was cut by EcoRI twice and Hindill only once. OB. The original piece of DNA was linear and was cut by EcoRI twice and Hindill only once. OC. The original piece of DNA was circular and was cut by EcoRI once and not at all by Hind!!!.arrow_forwardA result of agarose gel electrophoresis of plasmid DNA is shown below. The direction of the electrophoresis is indicated by the arrow head. Lane 1 shows the plasmid DNA with no restriction enzyme digestion. Lane 2 shows the plasmid DNA which has been digested by restriction enzyme ECORI. Given the fact that there are only relaxed and supercoiled forms existing in this plasmid, please annotate the bands in Lane 1 on this gel with their supercoiling status (supercoiled or relaxed) and explain why. Then based on the information from Lane 1 and Lane 2, analyze the number of EcoRI cleavage sites on this plasmid. Lane 1 2arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





